 (http://scientific.thomson.com/products/wos/)
(http://scientific.thomson.com/products/wos/)
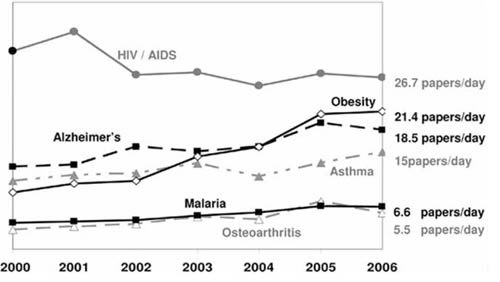
The number of research papers across both “popular” and less studied diseases currently far exceeds the reading time available to any one practitioner or analyst.
Specialised ontologies for well-defined scientific sub-domains across personal health are currently a time-saving rescue, and are developmentally aggressively pursued world-wide, both in the setting of ontology standards and in the actual building of the ontologies themselves. The pathway resources directory – PathGuide – alone, already references over 240 databases of biological pathways.
With exponential cost decreases in DNA sequencing during the last decade, personal genome sequencing, and therefore increasingly detailed and comprehensive genetic data is a special-case reality regarding the above, requiring urgent attention to the need for adequate information aggregators and filters across the many interrelated sub-domains of genomics-, molecular systems biology- and medical- (especially clinical and life-history) data.
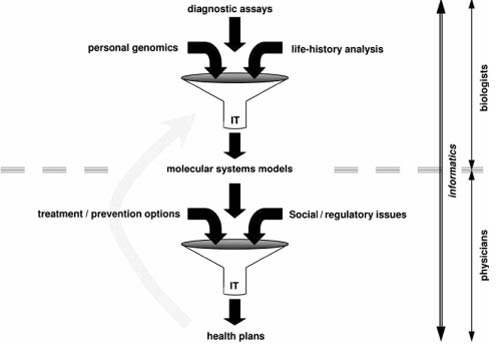
The large computational challenge is to find adequate semantic coupling between these compartmentalised and technically differentiated data repositories to produce useful aggregated output from the genotyping base up to predictive- and health plan levels.
With technical domain differentiation comes progress-stultifying terminology- and conceptual language complications:
 (ncbi.nlm.nih.gov/pmc/articles/PMC2674807/figure/F4/)
(ncbi.nlm.nih.gov/pmc/articles/PMC2674807/figure/F4/)
Gatfol is the technology that acts as coupling between
especially molecular biology and clinical IT environments
Gatfol’s patented multiword- to multiword semantic equivalence
transformation algorithms seamlessly “translate” between disparate
linguistic environments to provide instantaneous conceptual bridging
Computational Challenges of Personal Genomics
(Hamid Bolouri)
